Completed Projects
|
Finding improved agroecosystem management strategies that can continue to provide valuable economic products while improving ecosystem services are key to a complete portfolio for addressing climate change. Sustainable food production activities that actively sequester carbon are just one example. We investigated changes in carbon cycling on lands converted from conventional, row crop agriculture to management-intensive grazing (MiG) dairy systems in the southeast. These sandy soils, degraded from a long history of tillage and cotton cultivation were initially low in soil organic carbon (~0.6% C). Using a chronosequence approach we were able to show that soils were accumulating soil organic carbon at an extremely rapid rate, increasing cation exchange capacity and water holding capacity in the process. Carbon sequestration at these rates indicate this management style may increase soil health and serve as a net C sink for the first few years of implementation before returning to a net C source due to ruminant methane emissions. This work was conducted under the guidance of Aaron Thompson in collaboration with Megan Machmuller.
[Funding provided by the US Department of Agriculture’s, National Institute of Food and Agriculture (NIFA; Grant no. 2011-57003-30366) through the joint USDA-NASA-DOE Carbon Cycle Science Program.] |
|
|
|
Litter quality impacts the rate of microbial decomposition of organic matter (OM), with higher quality substrates leading to faster rates of decomposition. Since OM decomposition is the opposite of OM stabilization, one might presume higher quality substrates would lead to less OM stabilization. Yet at longer timescales, there is growing evidence higher quality substrates are used more efficiently and therefore contribute more to the mineral-associated organic fraction. For this project, I incubated soils with grass residues of differing maturity as well as manure analog of these same materials (ruminal digesta). This quality gradient allowed us to assess decomposition kinetics as well as monitor changes to the microbial community and silt and clay associated organic matter. Overall, we found that the decomposition of higher quality substrates led to larger amounts of clay-associated C and N.
[Funding provided by the US Department of Agriculture’s, National Institute of Food and Agriculture (NIFA; Grant no. 2011-57003-30366) through the joint USDA-NASA-DOE Carbon Cycle Science Program.] |
|
|
Before our AFRI-EWD project was funded, we first began by choosing a site location at Arnot Forest and hunting some soil microorganisms. Our goal was to use soil extract (solubilized, extractable soil organic matter - SESOM) derived from the organic rich surface soil (Oa horizon) as a culturing media for isolation. We created a library of soil isolates (bacterial and fungal) to then work with for future metabolomic studies. In the process, we found two new bacteria belonging to the genus Paraburkholderia, a genus of free-living as well as root-associated microorganisms that tend to have rather large genomes and exhibit high levels of metabolic diversity. We followed up with exometabolomic studies of one of these organisms (1N - now named Paraburkholderia solitsugae), growing in SESOM. We combined 1H NMR and LC-HRMS to track over 150 compounds in the media while also estimating carbon use efficiency during growth. Contrary to our expectations, we found no clear relationships between substrate nominal oxidation state (NOSC) or maximal growth rate (umax) and uptake preference (t50).
[Funding provided by: -AFRI Education and Workforce Development Program, grant no. 2019-67011-29513, from the US Department of Agriculture’s, National Institute of Food and Agriculture, Cornell University - CALS, Cornell University - CSBC NSF-IGERT, Atkinson Center for a Sustainable Future] |
Tracking the fate of low molecular weight organic substrates in solution-microbe-mineral systems
One way to begin unraveling the role of microbial metabolism in the formation of stable, mineral-associated organic
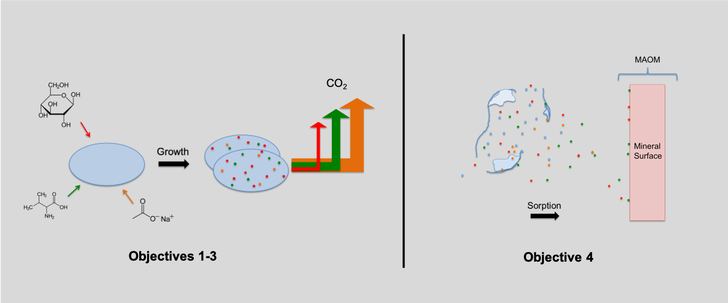
matter (MAOM) in soil is a model systems approach. This AFRI-EWD project sought to further our conceptual understanding of the processing of common, low molecular weight organic compounds in forest soil solution using a model systems approach. We utilized novel metabolomic methods (1H NMR, LC-HRMS) in the context of soil science in conjunction with stable isotope probing techniques (13C, 15N) to track substrate uptake kinetics and metabolic efficiency of use. We achieved these goals by: (1) - Assessing the usefulness of nominal oxidation state of carbon (NOSC) to predict microbial carbon use efficiency (CUE) of specific substrates in a mixed media (2) - Assessing the role of physiology on this relationship using isolates across a phylogenetic and growth rate gradient (3) - Determining the impact of substrate diversity on observed CUE (4) - Assessing if potentially divergent anabolic allocations of substrate carbon and nitrogen impact subsequent sorption affinity of substrate-derived biomass.
1 - Our results do not provide compelling population‐level support for NOSC as a predictive tool for either uptake kinetics or the efficiency of use of LMWOS in soil solution.
2 - Metabolic strategies of organisms are likely more important than chemical identity in determining LMWOS cycling in soils. Previous community‐level observations may be biased towards fast‐responding bacterial community members.
Check out the results from our first study in the Journal of Applied Microbiology - here
Impacts of substrate identity on sorption affinity of resulting necromass will soon be published! Stay tuned!
matter (MAOM) in soil is a model systems approach. This AFRI-EWD project sought to further our conceptual understanding of the processing of common, low molecular weight organic compounds in forest soil solution using a model systems approach. We utilized novel metabolomic methods (1H NMR, LC-HRMS) in the context of soil science in conjunction with stable isotope probing techniques (13C, 15N) to track substrate uptake kinetics and metabolic efficiency of use. We achieved these goals by: (1) - Assessing the usefulness of nominal oxidation state of carbon (NOSC) to predict microbial carbon use efficiency (CUE) of specific substrates in a mixed media (2) - Assessing the role of physiology on this relationship using isolates across a phylogenetic and growth rate gradient (3) - Determining the impact of substrate diversity on observed CUE (4) - Assessing if potentially divergent anabolic allocations of substrate carbon and nitrogen impact subsequent sorption affinity of substrate-derived biomass.
1 - Our results do not provide compelling population‐level support for NOSC as a predictive tool for either uptake kinetics or the efficiency of use of LMWOS in soil solution.
2 - Metabolic strategies of organisms are likely more important than chemical identity in determining LMWOS cycling in soils. Previous community‐level observations may be biased towards fast‐responding bacterial community members.
Check out the results from our first study in the Journal of Applied Microbiology - here
Impacts of substrate identity on sorption affinity of resulting necromass will soon be published! Stay tuned!
[Funding provided by the AFRI Education and Workforce Development Program, grant no. 2019-67011-29513, from the US Department of Agriculture’s, National Institute of Food and Agriculture.]
Unlocking the climate change mitigation potential of biochar composts
|
Co-led with fellow postdoc, Dr. Jialin Hu, this project used a laboratory incubation to test the impacts of biochar co-compost addition on emissions and soil health parameters. We assessed emissions trajectories following additions across a gradient of soil textures present in North Carolina. We utilized a mixture of GC-uECD and a modified LiCOR to measure N2O and CO2 emissions, respectively. We further measured N-speciation and enzyme activities through time to get a better mechanistic understanding of what processes were contributing to differences in N2O emissions.
|
|
Fingerprinting dissolved organic matter in rhizosphere soils
Fingerprinting dissolved organic matter in rhizosphere soils
|
|
Initially led by a post-doc in the Martinez Lab, Tapasya Babu, this project seeks to utilize metabolomic methods to better characterize changes in soil solution chemistry as affected by management (conventional vs. organic) as well as plant growth stage. To do this, we leveraged longterm research plots at Cornell's Musgrave Research Farm under different management practices. We re-created soil profiles in glass columns in the greenhouse with specially designed ports for soil solution sampling and inclusion of model clay fractions. Using the facilities at the Boyce Thompson Institute, we analyzed soil solution from the rhizosphere using LC-MS with switching polarity electrospray ionization. This technique, in conjunction with multiple available metabolomic data processing platforms (XCMS, MetaboAnalyst, MetaboQuest, ClassyFire) is enabling us to putatively identify thousands of molecules and focus on those that change across management, depth, and through time.
|
The relative importance of biotic and abiotic processes for understanding the dynamics of glyphosate herbicide in soils
Glyphosate [N-(phosphonomethyl)glycine] is the world’s most heavily applied herbicide. This project, led by Behrooz Azimzadeh, seeks to use a surface-enhanced approach to understand the contribution of biotic and abiotic degradation pathways on the removal of glyphosate in a model soil system. Though still in the initial stages, this work utilizes an isolate library created by undergraduate researcher, Manlin Jia, from soils collected at an upstate NY field site receiving glyphosate applications since 2014. Manlin used 4 different selective media to isolate bacteria that can grow on glyphosate as the sole source of respective elements (C, N, P, all). These isolates will be used in kinetic growth trials as well as flow-through ATR-FTIR experiments to better understand glyphosate biodegradation at surfaces. Experimental results can then be compared to abiotic, surface-catalyzed degradation by model Mn oxides.
[Funding provided in part by a Schmittau-Novak Grant from the School of Integrative Plant Science, Cornell University]